Graph Algorithms
Table of Contents
- Breadth First Traversal for a Graph
- Depth First Traversal for a Graph
- Dijkstra’s shortest path algorithm
- Floyd Warshall Algorithm (All Pairs Shortest Path)
- Union-Find Algorithm(Detect Cycle in an Undirected Graph)
- Kruskal’s Minimum Spanning Tree Algorithm
- Prim’s Minimum Spanning Tree
- Topological Sorting
- Articulation Points (or Cut Vertices) in a Graph
- Bridges in a graph
- Tarjan's strongly connected components algorithm
- Ford-Fulkerson Algorithm for Maximum Flow Problem
- Find minimum s-t cut in a flow network
- Dinic’s algorithm for Maximum Flow
- Check whether a given graph is Bipartite or not
- Maximum Bipartite Matching
- Hopcroft–Karp Algorithm for Maximum Matching
- cc
Breadth First Traversal for a Graph
#include <iostream> #include <list> using namespace std; class Graph { private: int v; list<int> *adj; public: Graph(int v); void AddEdge(int v, int w); /* print BFS from a given source s */ void BFS(int s); }; Graph::Graph(int v) { this->v = v; adj = new list<int>[v]; } void Graph::AddEdge(int v, int w) { adj[v].push_back(w); } void Graph::BFS(int s) { bool *visited = new bool[v]; for (int i = 0; i < v; ++i) { visited[i] = false; } list<int> queue; visited[s] = true; queue.push_back(s); list<int>::iterator it; int node; while (!queue.empty()) { node = queue.front(); cout << node << " "; queue.pop_front(); for (it = adj[node].begin(); it != adj[node].end(); ++it) { if (!visited[*it]) { visited[*it] = true; queue.push_back(*it); } } } cout << endl; delete[] visited; } int main() { Graph g(4); g.AddEdge(0, 1); g.AddEdge(0, 2); g.AddEdge(1, 2); g.AddEdge(2, 0); g.AddEdge(2, 3); g.AddEdge(3, 3); cout << "Following is Breadth First Traversal " << "(starting from vertex 2) \n"; g.BFS(2); return 0; }
Depth First Traversal for a Graph
#include <iostream> #include <list> using namespace std; class Graph { private: int v; list<int> *adj; void DFSHelper(int s, bool *visited); public: Graph(int v); void AddEdge(int v, int w); /* print DFS from a given source s */ void DFS(int s); }; Graph::Graph(int v) { this->v = v; adj = new list<int>[v]; } void Graph::AddEdge(int v, int w) { adj[v].push_back(w); } void Graph::DFSHelper(int s, bool *visited) { visited[s] = true; cout << s << " "; list<int>::iterator it; for (it = adj[s].begin(); it != adj[s].end(); ++it) { if (!visited[*it]) { DFSHelper(*it, visited); } } } void Graph::DFS(int s) { bool *visited = new bool[v]; for (int i = 0; i < v; ++i) { visited[i] = false; } DFSHelper(s, visited); } int main() { Graph g(4); g.AddEdge(0, 1); g.AddEdge(0, 2); g.AddEdge(1, 2); g.AddEdge(2, 0); g.AddEdge(2, 3); g.AddEdge(3, 3); cout << "Following is Depth First Traversal (starting from vertex 2) \n"; g.DFS(2); cout << endl; return 0; }
Dijkstra’s shortest path algorithm
- Create a set sptSet (shortest path tree set) that keeps track of vertices included in shortest path tree, i.e., whose minimum distance from source is calculated and finalized. Initially, this set is empty.
- Assign a distance value to all vertices in the input graph. Initialize all distance values as INFINITE. Assign distance value as 0 for the source vertex so that it is picked first.
- While sptSet doesn’t include all vertices a) Pick a vertex u which is not there in sptSetand has minimum distance value. b) Include u to sptSet. c) Update distance value of all adjacent vertices of u. To update the distance values, iterate through all adjacent vertices. For every adjacent vertex v, if sum of distance value of u (from source) and weight of edge u-v, is less than the distance value of v, then update the distance value of v.
#include <iostream> #include <cstdio> #include <climits> #define V 9 int MinDistance(int *dist, bool *spt_set) { int min = INT_MAX, min_index; for (int v = 0; v < V; ++v) { if (spt_set[v] == false && dist[v] < min) { min = dist[v]; min_index = v; } } return min_index; } void PrintSolution(int *dist) { printf("Vertex Distance from Source\n"); for (int i = 0; i < V; i++) { printf("%d \t\t %d\n", i, dist[i]); } } void dijkstra(int graph[V][V], int src) { int dist[V]; bool spt_set[V]; for (int i = 0; i < V; ++i) { dist[i] = INT_MAX; spt_set[i] = false; } dist[src] = 0; for (int count = 0; count < V - 1; ++count) { int min_dis_v = MinDistance(dist, spt_set); spt_set[min_dis_v] = true; for (int v = 0; v < V; ++v) { if (!spt_set[v] && graph[min_dis_v][v] && dist[min_dis_v] != INT_MAX && (dist[min_dis_v] + graph[min_dis_v][v] < dist[v])) { dist[v] = dist[min_dis_v] + graph[min_dis_v][v]; } } } PrintSolution(dist); } int main() { int graph[V][V] = {{0, 4, 0, 0, 0, 0, 0, 8, 0}, {4, 0, 8, 0, 0, 0, 0, 11, 0}, {0, 8, 0, 7, 0, 4, 0, 0, 2}, {0, 0, 7, 0, 9, 14, 0, 0, 0}, {0, 0, 0, 9, 0, 10, 0, 0, 0}, {0, 0, 4, 14, 10, 0, 2, 0, 0}, {0, 0, 0, 0, 0, 2, 0, 1, 6}, {8, 11, 0, 0, 0, 0, 1, 0, 7}, {0, 0, 2, 0, 0, 0, 6, 7, 0} }; dijkstra(graph, 0); return 0; }
Floyd Warshall Algorithm (All Pairs Shortest Path)
We initialize the solution matrix same as the input graph matrix as a first step. Then we update the solution matrix by considering all vertices as an intermediate vertex. The idea is to one by one pick all vertices and update all shortest paths which include the picked vertex as an intermediate vertex in the shortest path. When we pick vertex number k as an intermediate vertex, we already have considered vertices {0, 1, 2, .. k-1} as intermediate vertices. For every pair (i, j) of source and destination vertices respectively, there are two possible cases.
- k is not an intermediate vertex in shortest path from i to j. We keep the value of dist[i][j] as it is.
- k is an intermediate vertex in shortest path from i to j. We update the value of dist[i][j] as dist[i][k] + dist[k][j].
#include <cstdio> #include <climits> #define V 4 #define INF 9999 void PrintSolution(int dist[][V]) { printf ("Following matrix shows the shortest distances" " between every pair of vertices \n"); for (int i = 0; i < V; i++) { for (int j = 0; j < V; j++) { if (dist[i][j] == INF) printf("%7s", "INF"); else printf ("%7d", dist[i][j]); } printf("\n"); } } void FloydWarshall(int graph[][V]) { int dist[V][V], i, j , k; for (i = 0; i < V; ++i) { for (j = 0; j < V; ++j) { dist[i][j] = graph[i][j]; } } for (k = 0; k < V; ++k) { for (i = 0; i < V; ++i) { for (j = 0; j < V; ++j) { if (dist[i][k] + dist[k][j] < dist[i][j]) { dist[i][j] = dist[i][k] + dist[k][j]; } } } } PrintSolution(dist); } int main() { /* Let us create the following weighted graph 10 (0)------->(3) | /|\ 5 | | | | 1 \|/ | (1)------->(2) 3 */ int graph[V][V] = { {0, 5, INF, 10}, {INF, 0, 3, INF}, {INF, INF, 0, 1}, {INF, INF, INF, 0} }; FloydWarshall(graph); return 0; }
Union-Find Algorithm(Detect Cycle in an Undirected Graph)
- Find: Determine which subset a particular element is in. This can be used for determining if two elements are in the same subset.
- Union: Join two subsets into a single subset.
#include <cstdio> #include <cstdlib> #include <cstring> #define NO_PARENT -1 struct Edge { int src; int dest; }; struct Graph { // V-> Number of vertices int V; // E-> Number of edges int E; struct Edge* edge; }; struct Graph* CreateGraph(int V, int E) { struct Graph* graph = new Graph; graph->V = V; graph->E = E; graph->edge = new Edge[E]; return graph; } int Find(int *parent, int i) { if (parent[i] == NO_PARENT) { return i; } return Find(parent, parent[i]); } void Union(int *parent, int x, int y) { int xset = Find(parent, x); int yset = Find(parent, y); parent[xset] = yset; } int IsCycle(struct Graph* graph) { int *parent = new int[graph->V]; memset(parent, NO_PARENT, sizeof(int) * graph->V); for (int i = 0; i < graph->E; ++i) { int x = Find(parent, graph->edge[i].src); int y = Find(parent, graph->edge[i].dest); if (x == y) { return 1; } Union(parent, x, y); } delete[] parent; return 0; } int main() { /* Let us create following graph 0 | \ | \ 1-----2 */ int V = 3, E = 3; struct Graph* graph = CreateGraph(V, E); // add edge 0-1 graph->edge[0].src = 0; graph->edge[0].dest = 1; // add edge 1-2 graph->edge[1].src = 1; graph->edge[1].dest = 2; // add edge 0-2 graph->edge[2].src = 0; graph->edge[2].dest = 2; if (IsCycle(graph)) printf( "graph contains cycle\n" ); else printf( "graph doesn't contain cycle\n" ); delete graph; return 0; }
Union By Rank and Path Compression
The idea is to always attach smaller depth tree under the root of the deeper tree. This technique is called union by rank.
The second optimization to naive method is Path Compression. The idea is to flatten the tree when find() is called. When find() is called for an element x, root of the tree is returned. The find() operation traverses up from x to find root. The idea of path compression is to make the found root as parent of x so that we don’t have to traverse all intermediate nodes again.
The two techniques complement each other. The time complexity of each operations becomes even smaller than O(Logn).
./Files/union-by-rank-algorithm.cc
#include <cstdio> #include <cstdlib> #include <cstring> #define NO_PARENT -1 struct Edge { int src; int dest; }; struct Graph { // V-> Number of vertices int V; // E-> Number of edges int E; struct Edge* edge; }; struct Subset { int parent; int rank; }; struct Graph* CreateGraph(int V, int E) { struct Graph* graph = new Graph; graph->V = V; graph->E = E; graph->edge = new Edge[E]; return graph; } int Find(Subset *subsets, int i) { // find root and make root as parent of i (path compression) if (subsets[i].parent != i) { subsets[i].parent = Find(subsets, subsets[i].parent); } return subsets[i].parent; } void Union(Subset *subsets, int x, int y) { int xroot = Find(subsets, x); int yroot = Find(subsets, y); if (subsets[xroot].rank < subsets[yroot].rank) { subsets[xroot].parent = yroot; } else if (subsets[xroot].rank > subsets[yroot].rank) { subsets[yroot].parent = xroot; } else { subsets[yroot].parent = xroot; ++subsets[xroot].rank; } } int IsCycle(struct Graph* graph) { Subset *subsets = new Subset[graph->V]; for (int v = 0; v < graph->V; ++v) { subsets[v].parent = v; subsets[v].rank = 0; } for (int i = 0; i < graph->E; ++i) { int x = Find(subsets, graph->edge[i].src); int y = Find(subsets, graph->edge[i].dest); if (x == y) { return 1; } Union(subsets, x, y); } return 0; } int main() { /* Let us create following graph 0 | \ | \ 1-----2 */ int V = 3, E = 3; struct Graph* graph = CreateGraph(V, E); // add edge 0-1 graph->edge[0].src = 0; graph->edge[0].dest = 1; // add edge 1-2 graph->edge[1].src = 1; graph->edge[1].dest = 2; // add edge 0-2 graph->edge[2].src = 0; graph->edge[2].dest = 2; if (IsCycle(graph)) printf( "graph contains cycle\n" ); else printf( "graph doesn't contain cycle\n" ); delete graph; return 0; }
Kruskal’s Minimum Spanning Tree Algorithm
- Sort all the edges in non-decreasing order of their weight.
- Pick the smallest edge. Check if it forms a cycle with the spanning tree formed so far. If cycle is not formed, include this edge. Else, discard it.
- Repeat step#2 until there are (V-1) edges in the spanning tree.
#include <cstdio> #include <cstdlib> #include <cstring> struct Edge { int src; int dest; int weight; }; struct Graph { int V; int E; struct Edge *edge; }; struct Graph* CreateGraph(int V, int E) { struct Graph *graph = new Graph; graph->V = V; graph->E = E; graph->edge = new Edge[E]; return graph; } // A structure to represent a subset for union-find struct subset { int parent; int rank; }; int Find(struct subset *subsets, int i) { // find root and make root as parent of i (path compression) if (subsets[i].parent != i) { subsets[i].parent = Find(subsets, subsets[i].parent); } return subsets[i].parent; } // A function that does union of two sets of x and y // (uses union by rank) void Union(struct subset *subsets, int x, int y) { int xroot = Find(subsets, x); int yroot = Find(subsets, y); if (subsets[xroot].rank < subsets[yroot].rank) { subsets[xroot].parent = yroot; } else if (subsets[xroot].rank > subsets[yroot].rank) { subsets[yroot].parent = xroot; } else { // If ranks are same, then make one as root and increment // its rank by one subsets[yroot].parent = xroot; subsets[xroot].rank++; } } int EdgeComp(const void *a, const void *b) { struct Edge *e1 = (struct Edge*) a; struct Edge *e2 = (struct Edge*) b; return e1->weight > e2->weight; } void KruskalMST(struct Graph *graph) { qsort(graph->edge, graph->E, sizeof(graph->edge[0]), EdgeComp); struct subset *subsets = new subset[graph->V]; for (int v = 0; v < graph->V; ++v) { subsets[v].parent = v; subsets[v].rank = 0; } int e = 0; int i = 0; struct Edge result[graph->V]; while (e < graph->V - 1) { struct Edge next_edge = graph->edge[i++]; int x = Find(subsets, next_edge.src); int y = Find(subsets, next_edge.dest); if (x != y) { result[e++] = next_edge; Union(subsets, x, y); } // else discard the next_edge } printf("Following are the edges in the constructed MST\n"); for (i = 0; i < e; ++i) { printf("%d -- %d == %d\n", result[i].src, result[i].dest, result[i].weight); } delete[] subsets; } int main() { /* Let us create following weighted graph 10 0--------1 | \ | 6| 5\ |15 | \ | 2--------3 4 */ int V = 4; // Number of vertices in graph int E = 5; // Number of edges in graph struct Graph* graph = CreateGraph(V, E); // add edge 0-1 graph->edge[0].src = 0; graph->edge[0].dest = 1; graph->edge[0].weight = 10; // add edge 0-2 graph->edge[1].src = 0; graph->edge[1].dest = 2; graph->edge[1].weight = 6; // add edge 0-3 graph->edge[2].src = 0; graph->edge[2].dest = 3; graph->edge[2].weight = 5; // add edge 1-3 graph->edge[3].src = 1; graph->edge[3].dest = 3; graph->edge[3].weight = 15; // add edge 2-3 graph->edge[4].src = 2; graph->edge[4].dest = 3; graph->edge[4].weight = 4; KruskalMST(graph); delete[] graph->edge; delete graph; return 0; }
Prim’s Minimum Spanning Tree
- Create a set mstSet that keeps track of vertices already included in MST.
- Assign a key value to all vertices in the input graph. Initialize all key values as INFINITE. Assign key value as 0 for the first vertex so that it is picked first.
- While mstSet doesn’t include all vertices a) Pick a vertex u which is not there in mstSet and has minimum key value. b) Include u to mstSet. c) Update key value of all adjacent vertices of u. To update the key values, iterate through all adjacent vertices. For every adjacent vertex v, if weight of edge u-v is less than the previous key value of v, update the key value as weight of u-v
#include <cstdio> #include <climits> #define V 5 int MinKey(int *key, bool *mst_set) { int min = INT_MAX; int min_index; for (int v = 0; v < V; ++v) { if (mst_set[v] == false && key[v] < min) { min = key[v]; min_index = v; } } return min_index; } int PrintMST(int *parent, int graph[V][V]) { printf("Edge Weight\n"); for (int i = 1; i < V; i++) { printf("%d - %d %d \n", parent[i], i, graph[i][parent[i]]); } } void PrimMST(int graph[V][V]) { int key[V]; bool mst_set[V]; for (int i = 0; i < V; ++i) { key[i] = INT_MAX; mst_set[i] = false; } int parent[V]; key[0] = 0; parent[0] = -1; for (int count = 0; count < V - 1; ++count) { int min_v = MinKey(key, mst_set); mst_set[min_v] = true; for (int v = 0; v < V; ++v) { // graph[min_v][v] is non zero only for adjacent vertices of min_v // mstSet[v] is false for vertices not yet included in MST // Update the key only if graph[min_v][v] is smaller than key[v] if (graph[min_v][v] && mst_set[v] == false && graph[min_v][v] < key[v]) { parent[v] = min_v; key[v] = graph[min_v][v]; } } } PrintMST(parent, graph); } int main() { /* Let us create the following graph 2 3 (0)--(1)--(2) | / \ | 6| 8/ \5 |7 | / \ | (3)-------(4) 9 */ int graph[V][V] = {{0, 2, 0, 6, 0}, {2, 0, 3, 8, 5}, {0, 3, 0, 0, 7}, {6, 8, 0, 0, 9}, {0, 5, 7, 9, 0}, }; PrimMST(graph); return 0; }
Topological Sorting
Topological sorting for Directed Acyclic Graph (DAG) is a linear ordering of vertices such that for every directed edge uv, vertex u comes before v in the ordering. Topological Sorting for a graph is not possible if the graph is not a DAG.
#include<iostream> #include <list> #include <stack> using namespace std; class Graph { int V; list<int> *adj; void topologicalSortUtil(int v, bool visited[], stack<int> &Stack); public: Graph(int V); void addEdge(int v, int w); void topologicalSort(); }; Graph::Graph(int V) { this->V = V; adj = new list<int>[V]; } void Graph::addEdge(int v, int w) { adj[v].push_back(w); } void Graph::topologicalSortUtil(int v, bool visited[], stack<int> &Stack) { visited[v] = true; list<int>::iterator i; for (i = adj[v].begin(); i != adj[v].end(); ++i) if (!visited[*i]) topologicalSortUtil(*i, visited, Stack); Stack.push(v); } void Graph::topologicalSort() { stack<int> Stack; bool *visited = new bool[V]; for (int i = 0; i < V; i++) visited[i] = false; for (int i = 0; i < V; i++) if (visited[i] == false) topologicalSortUtil(i, visited, Stack); while (Stack.empty() == false) { cout << Stack.top() << " "; Stack.pop(); } cout << endl; } int main() { Graph g(6); g.addEdge(5, 2); g.addEdge(5, 0); g.addEdge(4, 0); g.addEdge(4, 1); g.addEdge(2, 3); g.addEdge(3, 1); cout << "Following is a Topological Sort of the given graph \n"; g.topologicalSort(); return 0; }
Articulation Points (or Cut Vertices) in a Graph
A O(V+E) algorithm to find all Articulation Points (APs) The idea is to use DFS (Depth First Search). In DFS, we follow vertices in tree form called DFS tree. In DFS tree, a vertex u is parent of another vertex v, if v is discovered by u (obviously v is an adjacent of u in graph). In DFS tree, a vertex u is articulation point if one of the following two conditions is true.
- u is root of DFS tree and it has at least two children.
- u is not root of DFS tree and it has a child v such that no vertex in subtree rooted with v has a back edge to one of the ancestors (in DFS tree) of u.
We do DFS traversal of given graph with additional code to find out
Articulation Points (APs). In DFS traversal, we maintain a parent[]
array where parent[u] stores parent of vertex u. Among the above
mentioned two cases, the first case is simple to detect. For every
vertex, count children. If currently visited vertex u is root
(parent[u] is NIL) and has more than two children, print it.
How to handle second case? The second case is trickier. We maintain an
array disc[] to store discovery time of vertices. For every node u, we
need to find out the earliest visited vertex (the vertex with minimum
discovery time) that can be reached from subtree rooted with u. So we
maintain an additional array low[] which is defined as follows.
low[u] = min(disc[u], disc[w])
where w is an ancestor of u and there is a back edge from
some descendant of u to w.
#include <iostream> #include <list> using namespace std; #define NIL -1 class Graph { public: Graph(int V); virtual ~Graph(); void AddEdge(int v, int w); void AP(); private: int V; list<int> *adj; void APUtil(int u, bool *visited, int *disc, int *low, int *parent, bool *ap); }; Graph::Graph(int V) { this->V = V; adj = new list<int>[V]; } Graph::~Graph() { delete[] adj; } void Graph::AddEdge(int v, int w) { adj[v].push_back(w); adj[w].push_back(v); // Note: the graph is undirected } void Graph::AP() { bool *visited = new bool[V]; int *disc = new int[V]; int *low = new int[V]; int *parent = new int[V]; bool *ap = new bool[V]; for (int i = 0; i < V; ++i) { parent[i] = NIL; visited[i] = false; ap[i] = false; } for (int i = 0; i < V; ++i) { if (visited[i] == false) { APUtil(i, visited, disc, low, parent, ap); } } for (int i = 0; i < V; ++i) { if (ap[i] == true) { cout << i << " "; } } delete[] visited; delete[] disc; delete[] low; delete[] parent; delete[] ap; } void Graph::APUtil(int u, bool *visited, int *disc, int *low, int *parent, bool *ap) { static int time = 0; int children = 0; visited[u] = true; disc[u] = low[u] = ++time; list<int>::iterator it; for (it = adj[u].begin(); it != adj[u].end(); ++it) { int v = *it; if (visited[v] == false) { children++; parent[v] = u; APUtil(v, visited, disc, low, parent, ap); low[u] = min(low[u], low[v]); // case 1 if (parent[u] == NIL && children > 1) { ap[u] = true; } // case 2 if (parent[u] != NIL && low[v] >= disc[u]) { ap[u] = true; } } else { if (v != parent[u]) { low[u] = min(low[u], disc[v]); } } } } int main() { // Create graphs given in above diagrams cout << "\nArticulation points in first graph \n"; Graph g1(5); g1.AddEdge(1, 0); g1.AddEdge(0, 2); g1.AddEdge(2, 1); g1.AddEdge(0, 3); g1.AddEdge(3, 4); g1.AP(); cout << "\nArticulation points in second graph \n"; Graph g2(4); g2.AddEdge(0, 1); g2.AddEdge(1, 2); g2.AddEdge(2, 3); g2.AP(); cout << "\nArticulation points in third graph \n"; Graph g3(7); g3.AddEdge(0, 1); g3.AddEdge(1, 2); g3.AddEdge(2, 0); g3.AddEdge(1, 3); g3.AddEdge(1, 4); g3.AddEdge(1, 6); g3.AddEdge(3, 5); g3.AddEdge(4, 5); g3.AP(); return 0; }
Bridges in a graph
An edge in an undirected connected graph is a bridge iff removing it disconnects the graph. For a disconnected undirected graph, definition is similar, a bridge is an edge removing which increases number of connected components.
A simple approach is to one by one remove all edges and see if removal of a edge causes disconnected graph. Following are steps of simple approach for connected graph.
- For every edge (u, v), do following a) Remove (u, v) from graph b) See if the graph remains connected (We can either use BFS or DFS) c) Add (u, v) back to the graph.
Time complexity of above method is O(E*(V+E)) for a graph represented using adjacency list.
A O(V+E) algorithm to find all Bridges
The idea is similar to O(V+E) algorithm for Articulation Points. We do
DFS traversal of the given graph. In DFS tree an edge(u, v) (u is
parent of v in DFS tree) is bridge if there does not exit any other
alternative to reach u or an ancestor of u from subtree rooted with v.
As discussed in the previous post, the value low[v] indicates earliest
visited vertex reachable from subtree rooted with v. The condition for
an edge(u, v) to be a bridge is, low[v] > disc[u].
#include <iostream> #include <list> using namespace std; #define NIL -1 class Graph { public: Graph(int V); virtual ~Graph(); void AddEdge(int v, int w); void Bridge(); private: int V; // No. of vertices list<int> *adj; void BridgeUtil(int v, bool visited[], int disc[], int low[], int parent[]); }; Graph::Graph(int V) { this->V = V; adj = new list<int>[V]; } Graph::~Graph() { delete[] adj; } void Graph::AddEdge(int v, int w) { adj[v].push_back(w); adj[w].push_back(v); // Note: the graph is undirected } void Graph::BridgeUtil(int u, bool visited[], int disc[], int low[], int parent[]) { static int time = 0; visited[u] = true; disc[u] = low[u] = ++time; list<int>::iterator it; for (it = adj[u].begin(); it != adj[u].end(); ++it) { int v = *it; if (!visited[v]) { parent[v] = u; BridgeUtil(v, visited, disc, low, parent); low[u] = min(low[u], low[v]); if (low[v] > disc[u]) { cout << u <<" " << v << endl; } } else if (v != parent[u]) { low[u] = min(low[u], disc[v]); } } } void Graph::Bridge() { bool *visited = new bool[V]; int *disc = new int[V]; int *low = new int[V]; int *parent = new int[V]; for (int i = 0; i < V; i++) { parent[i] = NIL; visited[i] = false; } for (int i = 0; i < V; i++) { if (visited[i] == false) { BridgeUtil(i, visited, disc, low, parent); } } delete[] visited; delete[] disc; delete[] low; delete[] parent; } int main() { cout << "\nBridges in first graph \n"; Graph g1(5); g1.AddEdge(1, 0); g1.AddEdge(0, 2); g1.AddEdge(2, 1); g1.AddEdge(0, 3); g1.AddEdge(3, 4); g1.Bridge(); cout << "\nBridges in second graph \n"; Graph g2(4); g2.AddEdge(0, 1); g2.AddEdge(1, 2); g2.AddEdge(2, 3); g2.Bridge(); cout << "\nBridges in third graph \n"; Graph g3(7); g3.AddEdge(0, 1); g3.AddEdge(1, 2); g3.AddEdge(2, 0); g3.AddEdge(1, 3); g3.AddEdge(1, 4); g3.AddEdge(1, 6); g3.AddEdge(3, 5); g3.AddEdge(4, 5); g3.Bridge(); return 0; }
Tarjan's strongly connected components algorithm
Overview
Tarjan's algorithm is an algorithm in graph theory for finding the strongly connected components of a graph. It runs in linear time, matching the time bound for alternative methods including Kosaraju's algorithm and the path-based strong component algorithm.
The algorithm in pseudocode
algorithm tarjan is input: graph G = (V, E) output: set of strongly connected components (sets of vertices) index := 0 S := empty for each v in V do if (v.index is undefined) then strongconnect(v) end if end for function strongconnect(v) // Set the depth index for v to the smallest unused index v.index := index v.lowlink := index index := index + 1 S.push(v) v.onStack := true // Consider successors of v for each (v, w) in E do if (w.index is undefined) then // Successor w has not yet been visited; recurse on it strongconnect(w) v.lowlink := min(v.lowlink, w.lowlink) else if (w.onStack) then // Successor w is in stack S and hence in the current SCC v.lowlink := min(v.lowlink, w.index) end if end for // If v is a root node, pop the stack and generate an SCC if (v.lowlink = v.index) then start a new strongly connected component repeat w := S.pop() w.onStack := false add w to current strongly connected component while (w != v) output the current strongly connected component end if end function
Strong connectivity
In undirected graphs, two vertices are connected if they have a path connecting them. How should we define connected in a directed graph?
We say that a vertex a is strongly connected to b if there exist two paths, one from a to b and another from b to a.
Note that we allow the two paths to share vertices or even to share edges. We will use a ~ b as shorthand for "a is strongly connected to b". We will allow very short paths, with one vertex and no edges, so that any vertex is strongly connected to itself.
Strong connectivity algorithm1
Define the DFS numbering dfsnum(v) to be the number of vertices visited before v in the DFS. Then if there is a back or cross edge out of the subtree of v, it's to something visited before v and therefore with a smaller dfsnum. We use this by defining the low value low(v) to be the smallest dfsnum of a vertex reachable by a back or cross edge from the subtree of v. If there is no such edge, low(v)=dfsnum(v). Then rephrasing what we've seen so far, v is a head of a component exactly when low(v)=dfsnum(v). The advantage of using these definitions is that dfsnum(v) is trivial to calculate as we perform the DFS, and low(v) is easily computed by combining the low values from the children of v with the values coming from back or cross edges out of v itself.
We use one more simple data structure, a stack L (represented as a list) which we use to identify the subtree rooted at a vertex. We simply push each new vertex onto L as we visit it; then when we have finished visiting a vertex, its subtree will be everything pushed after it onto L. If v is a head, and we've already deleted the other heads in that subtree, the remaining vertices left on L will be exactly the component [v].
We are now ready to describe the actual algorithm. It simply performs a DFS, keeping track of the low and dfsnum values defined above, using them to identify heads of components, and when finding a head deleting the whole component from the graph, using L to find the vertices of the component.
Algorithm in Python
def strongly_connected_components(graph): """ Tarjan's Algorithm (named for its discoverer, Robert Tarjan) is a graph theory algorithm for finding the strongly connected components of a graph. Based on: http://en.wikipedia.org/wiki/Tarjan%27s_strongly_connected_components_algorithm """ index_counter = [0] stack = [] lowlinks = {} index = {} result = [] def strongconnect(node): # set the depth index for this node to the smallest unused index index[node] = index_counter[0] lowlinks[node] = index_counter[0] index_counter[0] += 1 stack.append(node) # Consider successors of `node` try: successors = graph[node] except: successors = [] for successor in successors: if successor not in lowlinks: # Successor has not yet been visited; recurse on it strongconnect(successor) lowlinks[node] = min(lowlinks[node],lowlinks[successor]) elif successor in stack: # the successor is in the stack and hence in the current strongly connected component (SCC) lowlinks[node] = min(lowlinks[node],index[successor]) # If `node` is a root node, pop the stack and generate an SCC if lowlinks[node] == index[node]: connected_component = [] while True: successor = stack.pop() connected_component.append(successor) if successor == node: break component = tuple(connected_component) # storing the result result.append(component) for node in graph: if node not in lowlinks: strongconnect(node) return result if __name__ == '__main__': result = strongly_connected_components( {1:[2],2:[1,5],3:[4],4:[3,5],5:[6],6:[7],7:[8],8:[6,9],9:[]} ); print result
Ford-Fulkerson Algorithm for Maximum Flow Problem
Given a graph which represents a flow network where every edge has a capacity. Also given two vertices source ‘s’ and sink ‘t’ in the graph, find the maximum possible flow from s to t with following constraints:
- Flow on an edge doesn’t exceed the given capacity of the edge.
- Incoming flow is equal to outgoing flow for every vertex except s and t.
For example, consider the following graph.
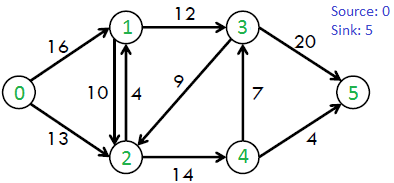
The maximum possible flow in the above graph is 23.
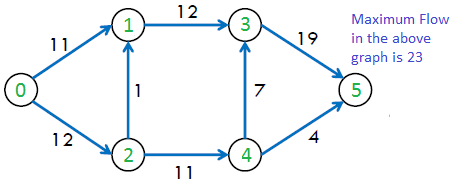
Residual Graph of a flow network is a graph which indicates additional possible flow. If there is a path from source to sink in residual graph, then it is possible to add flow. Every edge of a residual graph has a value called residual capacity which is equal to original capacity of the edge minus current flow. Residual capacity is basically the current capacity of the edge.
The important thing is, we need to update residual capacities in the residual graph. We subtract path flow from all edges along the path and we add path flow along the reverse edges We need to add path flow along reverse edges because may later need to send flow in reverse direction.
The above implementation of Ford Fulkerson Algorithm is called Edmonds-Karp Algorithm. The idea of Edmonds-Karp is to use BFS in Ford Fulkerson implementation as BFS always picks a path with minimum number of edges. When BFS is used, the worst case time complexity can be reduced to O(VE2). The above implementation uses adjacency matrix representation though where BFS takes O(V2) time, the time complexity of the above implementation is O(EV3)
#include <iostream> #include <queue> #include <climits> #include <cstring> using namespace std; #define V 6 bool BFS(int graph[V][V], int s, int t, int parent[]) { bool visited[V]; memset(visited, 0 , sizeof(visited)); queue<int> q; q.push(s); visited[s] = true; parent[s] = -1; while (!q.empty()) { int u = q.front(); q.pop(); for (int v = 0; v < V; ++v) { if (visited[v] == false && graph[u][v] > 0) { q.push(v); parent[v] = u; visited[v] = true; } } } return visited[t] == true; } int FordFulkerson(int graph[V][V], int s, int t) { int u, v; int resi_graph[V][V]; for (u = 0; u < V; ++u) { for (v = 0; v < V; ++v) { resi_graph[u][v] = graph[u][v]; } } int parent[V]; int max_flow = 0; while (BFS(resi_graph, s, t, parent)) { int path_flow = INT_MAX; for (v = t; v != s; v = parent[v]) { u = parent[v]; path_flow = min(path_flow, resi_graph[u][v]); } for (v = t; v != s; v = parent[v]) { u = parent[v]; resi_graph[u][v] -= path_flow; resi_graph[v][u] += path_flow; } max_flow += path_flow; } return max_flow; } int main() { int graph[V][V] = { {0, 16, 13, 0, 0, 0}, {0, 0, 10, 12, 0, 0}, {0, 4, 0, 0, 14, 0}, {0, 0, 9, 0, 0, 20}, {0, 0, 0, 7, 0, 4}, {0, 0, 0, 0, 0, 0} }; cout << "The maximum possible flow from 0 to 5 is " << FordFulkerson(graph, 0 ,5) << endl; return 0; }
Find minimum s-t cut in a flow network
In a flow network, an s-t cut is a cut that requires the source ‘s’ and the sink ‘t’ to be in different subsets, and it consists of edges going from the source’s side to the sink’s side. The capacity of an s-t cut is defined by the sum of capacity of each edge in the cut-set.2
The problem discussed here is to find minimum capacity s-t cut of the given network. Expected output is all edges of the minimum cut.
For example, in the following flow network, example s-t cuts are {{0 ,1}, {0, 2}}, {{0, 2}, {1, 2}, {1, 3}}, etc. The minimum s-t cut is {{1, 3}, {4, 3}, {4 5}} which has capacity as 12+7+4 = 23.
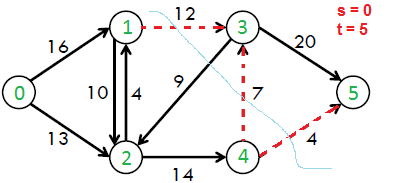
Like Maximum Bipartite Matching, this is another problem which can solved using Ford-Fulkerson Algorithm. This is based on max-flow min-cut theorem.
The max-flow min-cut theorem states that in a flow network, the amount of maximum flow is equal to capacity of the minimum cut. See CLRS book for proof of this theorem.
From Ford-Fulkerson, we get capacity of minimum cut. How to print all edges that form the minimum cut? The idea is to use residual graph.
Following are steps to print all edges of minimum cut.
- Run Ford-Fulkerson algorithm and consider the final residual graph.
- Find the set of vertices that are reachable from source in the residual graph.
- All edges which are from a reachable vertex to non-reachable vertex are minimum cut edges. Print all such edges.
#include <cstring> #include <iostream> #include <queue> #include <limits> using namespace std; #define V 6 /* Returns true if there is a path from source 's' to sink 'd' in residual graph. Also fills parent[] to store the path */ bool Bfs(int res_graph[V][V], int s, int d, int *parent) { bool visited[V]; memset(visited, 0, sizeof(visited)); queue<int> q; q.push(s); visited[s] = true; parent[s] = -1; while (!q.empty()) { int u = q.front(); q.pop(); for (int v = 0; v < V; ++v) { if (visited[v] == false && res_graph[u][v] > 0) { q.push(v); parent[v] = u; visited[v] = true; } } } return (visited[d] == true); } void Dfs(int res_graph[V][V], int s, bool *visited) { visited[s] = true; for (int i = 0; i < V; ++i) { if (res_graph[s][i] && !visited[i]) { Dfs(res_graph, i, visited); } } } void MinCut(int graph[V][V], int s, int d) { int u, v; int res_graph[V][V]; for (u = 0; u < V; ++u) { for (v = 0; v < V; ++v) { res_graph[u][v] = graph[u][v]; } } int parent[V]; while (Bfs(res_graph, s, d, parent)) { int path_flow = numeric_limits<int>::max(); for (v = d; v != s; v = parent[v]) { u = parent[v]; path_flow = min(path_flow, res_graph[u][v]); } for (v = d; v != s; v = parent[v]) { u = parent[v]; res_graph[u][v] -= path_flow; res_graph[v][u] += path_flow; } } bool visited[V]; memset(visited, false, sizeof(visited)); Dfs(res_graph, s, visited); for (int i = 0; i < V; ++i) { for (int j = 0; j < V; ++j) { if (visited[i] && !visited[j] && graph[i][j]) { cout << i << " - " << j << endl; } } } } int main() { int graph[V][V] = { {0, 16, 13, 0, 0, 0}, {0, 0, 10, 12, 0, 0}, {0, 4, 0, 0, 14, 0}, {0, 0, 9, 0, 0, 20}, {0, 0, 0, 7, 0, 4}, {0, 0, 0, 0, 0, 0} }; MinCut(graph, 0, 5); return 0; }
Dinic’s algorithm for Maximum Flow
Time complexity of Edmond Karp Implementation is O(VE2). In this post, a new Dinic’s algorithm is discussed which is a faster algorithm and takes O(EV2).
Like Edmond Karp’s algorithm, Dinic’s algorithm uses following concepts
- A flow is maximum if there is no s to t path in residual graph.
- BFS is used in a loop. There is a difference though in the way we use BFS in both algorithms.
In Edmond’s Karp algorithm, we use BFS to find an augmenting path and send flow across this path. In Dinic’s algorithm, we use BFS to check if more flow is possible and to construct level graph. In level graph, we assign levels to all nodes, level of a node is shortest distance (in terms of number of edges) of the node from source. Once level graph is constructed, we send multiple flows using this level graph. This is the reason it works better than Edmond Karp. In Edmond Karp, we send only flow that is send across the path found by BFS.
Dinic’s algorithm:
1) Initialize residual graph G as given graph.
1) Do BFS of G to construct a level graph (or
assign levels to vertices) and also check if
more flow is possible.
a) If more flow is not possible, then return.
b) Send multiple flows in G using level graph
until blocking flow is reached. Here using
level graph means, in every flow,
levels of path nodes should be 0, 1, 2...
(in order) from s to t.
+ Illustration of Dinic’s algorithm in the WiKi.
- And more: http://e-maxx.ru/algo/dinic
#include <iostream> #include <vector> #include <list> #include <limits> using namespace std; struct Edge { int v; int flow; int c; }; class Graph { public: Graph(int V); void AddEdge(int u, int v, int c); bool BFS(int s, int t); int SendFlow(int u, int flow, int t, int *start); int DinicMaxFlow(int s, int t); private: int V; int *level; vector<Edge> *adj; }; Graph::Graph(int V) { adj = new vector<Edge>[V]; this->V = V; level = new int[V]; } void Graph::AddEdge(int u, int v, int c) { // Forward edge : 0 flow and c capacity Edge a{v, 0, c}; // Back edge : 0 flow and 0 capacity Edge b{u, 0, 0}; adj[u].push_back(a); adj[v].push_back(b); } bool Graph::BFS(int s, int t) { for (int i = 0; i < V; ++i) { level[i] = -1; } level[s] = 0; list<int> q; q.push_back(s); vector<Edge>::iterator it; while (!q.empty()) { int u = q.front(); q.pop_front(); for (it = adj[u].begin(); it != adj[u].end(); ++it) { Edge &e = *it; if (level[e.v] < 0 && e.flow < e.c) { level[e.v] = level[u] + 1; q.push_back(e.v); } } } // IF we can not reach to the sink we // return false else true return level[t] < 0 ? false : true; } // A DFS based function to send flow after BFS has // figured out that there is a possible flow and // constructed levels. This function called multiple // times for a single call of BFS. // flow: Current flow send by parent function call // start[]: To keep track of next edge to be explored. // start[i] stores count of edges explored // from i. // u: Current vertex // t: Sink int Graph::SendFlow(int u, int flow, int t, int *start) { if (u == t) { return flow; } for (; start[u] < adj[u].size(); start[u]++) { Edge &e = adj[u][start[u]]; if (level[e.v] == level[u]+1 && e.flow < e.c) { int curr_flow = min(flow, e.c - e.flow); int temp_flow = SendFlow(e.v, curr_flow, t, start); if (temp_flow > 0) { // add flow to current edge e.flow += temp_flow; return temp_flow; } } } return 0; } int Graph::DinicMaxFlow(int s, int t) { if (s == t) { return -1; } int total = 0; while (BFS(s , t) == true) { int *start = new int[V+1]; while (int flow = SendFlow(s, numeric_limits<int>::max(), t, start)) { total += flow; } } return total; } int main() { Graph g(6); g.AddEdge(0, 1, 16 ); g.AddEdge(0, 2, 13 ); g.AddEdge(1, 2, 10 ); g.AddEdge(1, 3, 12 ); g.AddEdge(2, 1, 4 ); g.AddEdge(2, 4, 14); g.AddEdge(3, 2, 9 ); g.AddEdge(3, 5, 20 ); g.AddEdge(4, 3, 7 ); g.AddEdge(4, 5, 4); cout << "Maximum flow " << g.DinicMaxFlow(0, 5) << endl; return 0; }
Check whether a given graph is Bipartite or not
A Bipartite Graph is a graph whose vertices can be divided into two independent sets, U and V such that every edge (u, v) either connects a vertex from U to V or a vertex from V to U. In other words, for every edge (u, v), either u belongs to U and v to V, or u belongs to V and v to U. We can also say that there is no edge that connects vertices of same set.
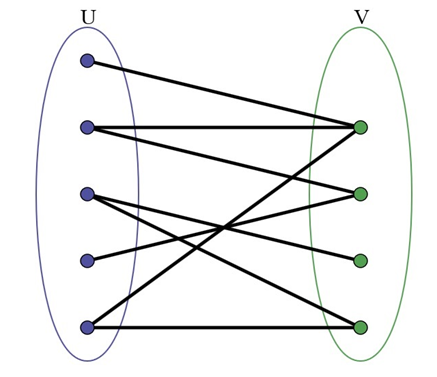
A bipartite graph is possible if the graph coloring is possible using two colors such that vertices in a set are colored with the same color.
Following is a simple algorithm to find out whether a given graph is Birpartite or not using Breadth First Search (BFS).
- Assign RED color to the source vertex (putting into set U).
- Color all the neighbors with BLUE color (putting into set V).
- While assigning colors, if we find a neighbor which is colored with same color as current vertex, then the graph cannot be colored with 2 vertices (or graph is not Bipartite)
#include <iostream> #include <queue> using namespace std; #define V 4 bool IsBipartiteUtil(int G[][V], int src, int *color_arr) { color_arr[src] = 1; queue<int> q; q.push(src); while (!q.empty()) { int u = q.front(); q.pop(); for (int v = 0; v < V; ++v) { if (G[u][v] && color_arr[v] == -1) { color_arr[v] = 1 - color_arr[u]; q.push(v); } else if (G[u][v] && color_arr[v] == color_arr[u]) { return false; } } } return true; } bool IsBipartite(int G[][V]) { // Create a color array to store colors assigned to all veritces. int color_arr[V]; for (int i = 0; i < V; ++i) { color_arr[i] = -1; } // This code is to handle disconnected graoh for (int i = 0; i < V; ++i) { if (color_arr[i] == -1) { if (IsBipartiteUtil(G, i, color_arr) == false) { return false; } } } return true; } int main() { int G[][V] = {{0, 1, 0, 1}, {1, 0, 1, 0}, {0, 1, 0, 1}, {1, 0, 1, 0}}; IsBipartite(G) ? cout << "Yes" : cout << "No"; return 0; }
Maximum Bipartite Matching
A matching in a Bipartite Graph is a set of the edges chosen in such a way that no two edges share an endpoint. A maximum matching is a matching of maximum size (maximum number of edges). In a maximum matching, if any edge is added to it, it is no longer a matching. There can be more than one maximum matchings for a given Bipartite Graph.
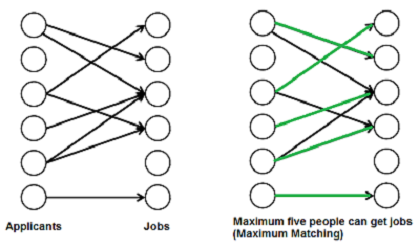
In bpm(), we one by one try all jobs that an applicant ‘u’ is interested in until we find a job, or all jobs are tried without luck. For every job we try, we do following.
If a job is not assigned to anybody, we simply assign it to the applicant and return true. If a job is assigned to somebody else say x, then we recursively check whether x can be assigned some other job. To make sure that x doesn’t get the same job again, we mark the job ‘v’ as seen before we make recursive call for x. If x can get other job, we change the applicant for job ‘v’ and return true.
./Files/max_bipartite_matching.cc
#include <cstring> #include <iostream> using namespace std; #define M 6 #define N 6 // A DFS based recursive function that returns true if a // matching for vertex u is possible bool Bpm(bool bp_graph[M][N], int u, bool *seen, int *match_r) { for (int v = 0; v < N; ++v) { if (bp_graph[u][v] && !seen[v]) { seen[v] = true; // If job 'v' is not assigned to an applicant OR // previously assigned applicant for job v (which is matchR[v]) // has an alternate job available. if (match_r[v] < 0 || Bpm(bp_graph, match_r[v], seen, match_r)) { match_r[v] = u; return true; } } } return false; } int MaxBPM(bool bp_graph[M][N]) { // An array to keep track of the applicants assigned to jobs. int match_r[N]; memset(match_r, -1, sizeof(match_r)); int result = 0; for (int u = 0; u < M; ++u) { bool seen[N]; memset(seen, 0, sizeof(seen)); if (Bpm(bp_graph, u, seen, match_r)) { ++result; } } return result; } int main() { bool bp_graph[M][N] = {{0, 1, 1, 0, 0, 0}, {1, 0, 0, 1, 0, 0}, {0, 0, 1, 0, 0, 0}, {0, 0, 1, 1, 0, 0}, {0, 0, 0, 0, 0, 0}, {0, 0, 0, 0, 0, 1}}; cout << "Maximum maching is " << MaxBPM(bp_graph) << endl; return 0; }
Hopcroft–Karp Algorithm for Maximum Matching
Time complexity of the Ford Fulkerson based algorithm is O(V x E). Hopcroft Karp algorithm is an improvement that runs in O(√V x E) time.
The Hopcroft Karp algorithm is based on below concept.
A matching M is not maximum if there exists an augmenting path. It is also true other way, i.e, a matching is maximum if no augmenting path exists
The idea is to use BFS (Breadth First Search) to find augmenting paths. Since BFS traverses level by level, it is used to divide the graph in layers of matching and not matching edges. A dummy vertex NIL is added that is connected to all vertices on left side and all vertices on right side. Following arrays are used to find augmenting path. Distance to NIL is initialized as INF (infinite). If we start from dummy vertex and come back to it using alternating path of distinct vertices, then there is an augmenting path.
Once an augmenting path is found, DFS (Depth First Search) is used to add augmenting paths to current matching. DFS simply follows the distance array setup by BFS. It fills values in pairU[u] and pairV[v] if v is next to u in BFS.
The below implementation is mainly adopted from the algorithm provided on Wiki page of Hopcroft Karp algorithm.
./Files/hopcroft-karp_for_max_matching.cc
#include <climits> #include <iostream> #include <list> #include <queue> using namespace std; #define NIL 0 #define INF INT_MAX class BipGraph { public: BipGraph(int m, int n); void AddEdge(int u, int v); // Returns true if there is an augmenting path bool Bfs(); // Adds augmenting path if there is one beginning with u bool Dfs(int u); // Returns size of maximum matcing int HopcroftKarp(); private: // m and n are number of vertices on left // and right sides of Bipartite Graph int m, n; list<int> *adj; int *pair_u, *pair_v, *dist; }; BipGraph::BipGraph(int m, int n) { this->m = m; this->n = n; adj = new list<int>[m+1]; } void BipGraph::AddEdge(int u, int v) { adj[u].push_back(v); adj[v].push_back(u); } int BipGraph::HopcroftKarp() { pair_u = new int[m+1]; pair_v = new int[n+1]; // dist[u] stores distance of left side vertices // dist[u] is one more than dist[u'] if u is next // to u'in augmenting path dist = new int[m+1]; for (int u = 0; u <= m; ++u) { pair_u[u] = NIL; } for (int v = 0; v <= n; ++v) { pair_v[v] = NIL; } int result = 0; // Keep updating the result while there is an // augmenting path. while (Bfs()) { // Find a free vertex for (int u = 1; u <= m; ++u) { // If current vertex is free and there is // an augmenting path from current vertex if (pair_u[u] == NIL && Dfs(u)) { ++result; } } } return result; } bool BipGraph::Bfs() { queue<int> que; for (int u = 1; u <= m; ++u) { if (pair_u[u] == NIL) { // If this is a free vertex, add it to queue dist[u] = 0; que.push(u); } else { // Else set distance as infinite so that this vertex // is considered next time dist[u] = INF; } } dist[NIL] = INF; while (!que.empty()) { int u = que.front(); que.pop(); if (dist[u] < dist[NIL]) { // If this node is not NIL and can provide a shorter path to NIL list<int>::iterator it; for (it = adj[u].begin(); it != adj[u].end(); ++it) { int v = *it; if (dist[pair_v[v]] == INF) { dist[pair_v[v]] = dist[u] + 1; que.push(pair_v[v]); } } } } return (dist[NIL] != INF); } bool BipGraph::Dfs(int u) { if (u != NIL) { list<int>::iterator it; for (it = adj[u].begin(); it != adj[u].end(); ++it) { int v = *it; if (dist[pair_v[v]] == dist[u] + 1) { if (Dfs(pair_v[v]) == true) { pair_v[v] = u; pair_u[u] = v; return true; } } } // If there is no augmenting path beginning with u. dist[u] = INF; return false; } return true; } int main() { BipGraph g(4, 4); g.AddEdge(1, 2); g.AddEdge(1, 3); g.AddEdge(2, 1); g.AddEdge(3, 2); g.AddEdge(4, 2); g.AddEdge(4, 4); cout << "Size of maximum matching is " << g.HopcroftKarp(); return 0; }
cc
INCLUDE: "./Files/" src c+